Projectes de recerca oberts a candidatures
Research projects
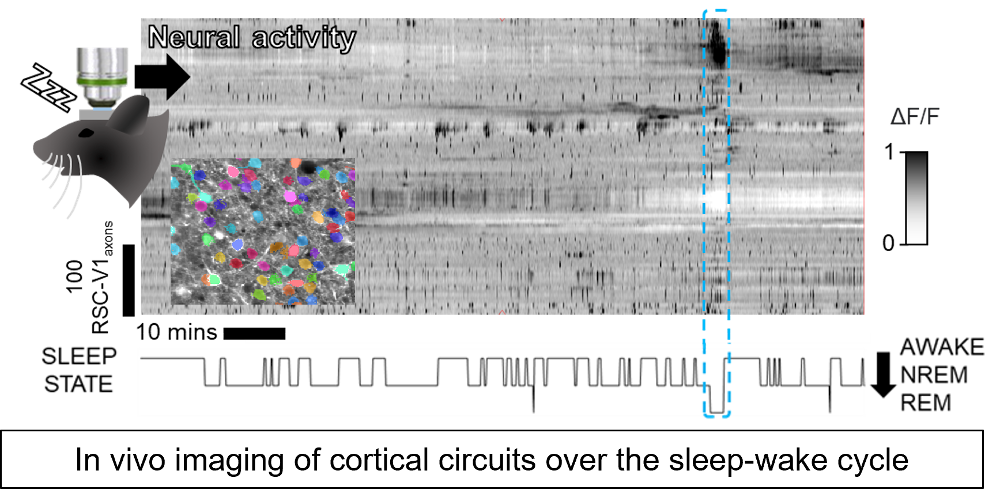
During sleep and dreaming the brain internally produces realistic simulations which are thought to play an important role in learning, creativity and memory. The project aims to gain a mechanistic understanding of the circuits through which the brain can internally generate this highly structured cortical neural activity during sleep, how this activity is coordinated between brain regions, and the function it serves in learning, creativity and memory. To answer these questions, we use in vivo brain imaging to probe and manipulate cortical circuits at cellular resolution in in mice during learning and sleep.
The candidate will have access to state-of-the-art in vivo 2-photon calcium imaging, holographic optogenetic and virtual reality approaches to measure and perturb neural circuits in awake behaving and sleeping mice at cellular and subcellular resolution.
Candidates, please contact Dr. Adam Ranson: adam.ranson@uab.cat
In your application please include:
- A CV describing relevant experience and skills. This should include any practical experimental experience and knowledge of data analysis approaches (such as in python or MATLAB).
- An expression of interest letter describing why you are attracted to this position.
- In the case of applications to PhD fellowships, full undergraduate transcripts and a document summarising overall GPA for undergraduate and masters courses. Ideally these should be provided as converted to the Spanish system using the following website.
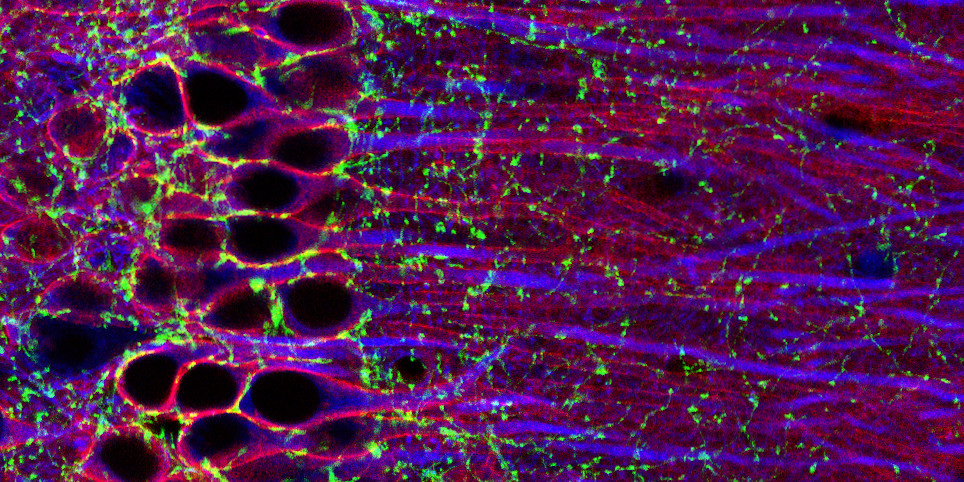
The project aims to gain a mechanistic understanding of how cannabinoids such as THC and CBD act in the brain—which circuits and cell types are engaged, what molecular programs are recruited, and how these mechanisms relate to behavior. To address these questions, we will combine snRNA-seq, translatomics, and lipidomics with mouse behavioral studies and targeted experimental perturbations.
The candidate will have access to state-of-the-art omics platforms and in vivo mouse behavioral approaches, and will work within a highly collaborative environment with expertise in cannabinoid neuropharmacology, circuit neuroscience, and translational models relevant to CNS disease.
Applicants should have a strong background in neuroscience/neuropharmacology and an interest in how cannabinoids modulate brain circuits and behavior. Experience in at least one of the following is expected: -Omics: snRNA-seq, translatomics, and/or lipidomics; -Circuit methods (e.g., stereotaxic surgery, viral tools) and mouse behavior; -Data analysis in R/Python for high-dimensional datasets.
Candidates, please contact Dr. Emma Puighermanal: emma.puighermanal@uab.cat
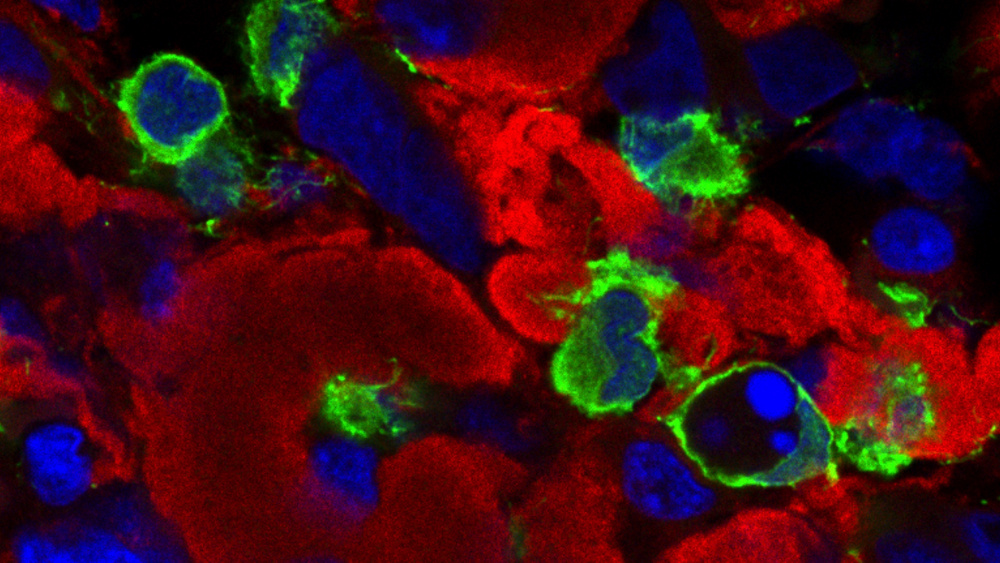
Our research is focused on exploring the different microenvironments of human glioma samples, and analyze experimental models in vivo and in vitro, to understand the function of immune cells, especially microglia-macrophages to achieve tumor clearance. The outcome of this project will suggest novel strategies for immunotherapy.
Techniques: Mapping the cellular populations in tissue from biopsies of patients and glioblastoma models in mice by using high-resolution immunohistofluorescence and histocytometry. Use in vivo and in vitro models to study and test immunotherapeutic strategies.
Candidates should have knowledge on advanced microscopy (Confocal Imaging). Previous experience with animal experimentation and particularly, stereotaxic surgery in mouse brain will be highly valued.
Candidates, please contact Dr. Carlos Barcia: carlos.barcia@uab.cat
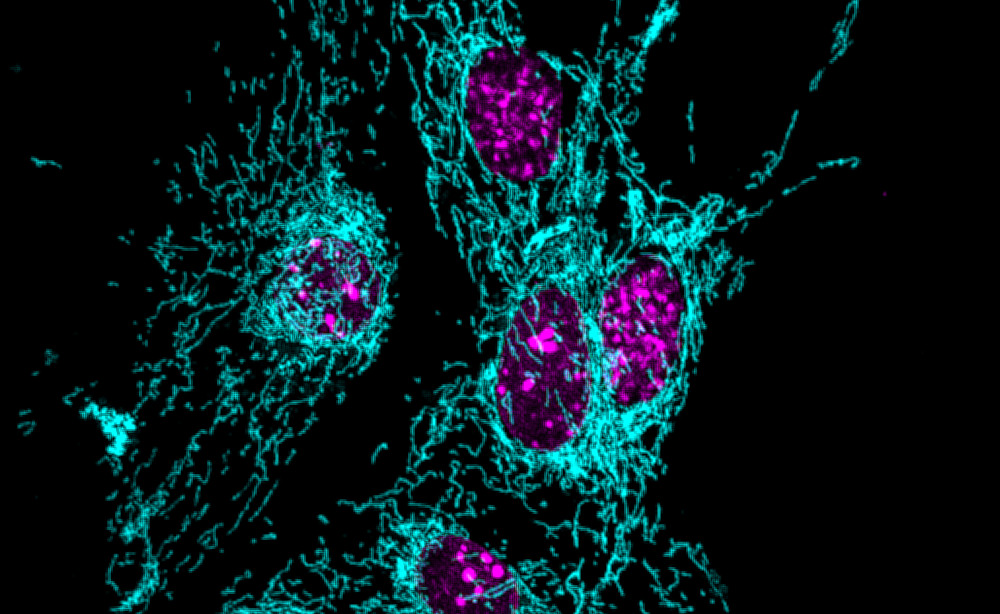
The project aims to uncover the molecular mechanisms that drive selective neuronal vulnerability in primary mitochondrial disease, focusing on how defects in mitochondrial complex I trigger neurodegeneration. To address this, we will use a Ndufs4-deficient mouse model and combine transcriptional, metabolic, and biochemical profiling with high-resolution imaging, pharmacological interventions, and behavioral analyses to dissect PKR-mediated antiviral responses, lipid droplet dynamics, and their contribution to neuronal death.
The candidate will have access to cutting-edge omics platforms and in vivo mouse behavioral assays, and will be embedded in a highly collaborative research environment.
Applicants should have a strong background in neuroscience, cell biology, or a related field, and a clear interest in the molecular and cellular mechanisms underlying neurodegeneration and mitochondrial disease. Experience in at least one of the following areas is expected: 1) Omics approaches, such as transcriptomics, metabolomics, and/or lipidomics; 2) In vivo mouse work, including disease models, pharmacological interventions, and histological or imaging techniques; 3) Data analysis using R and/or Python, particularly for high-dimensional biological datasets.
Candidates, please contact Dr. Albert Quintana: albert.quintana@uab.cat
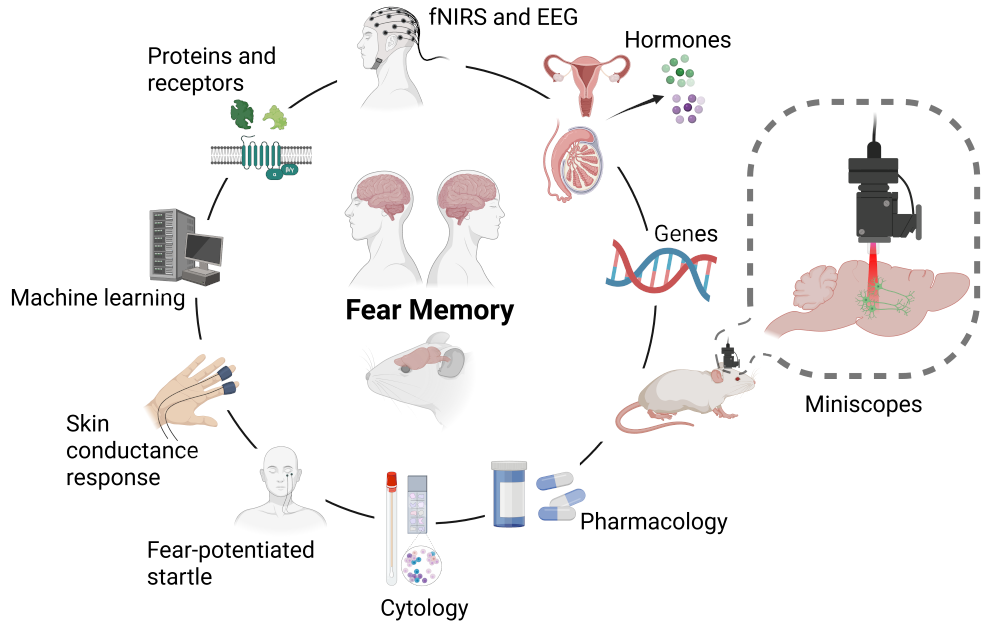
We aim to deepen our understanding of how stress alters memory networks in the brain. Our multidisciplinary team conducts research in both rodents and humans, using machine learning to integrate findings across species. Examples of recent publications:
- Stress-induced changes in the molecular processes underlying fear memories: implications for PTSD and relevant animal models
- Sex differences in neural projections of fear memory processing in mice and humans
- PACAP-PAC1R modulates fear extinction via the ventromedial hypothalamus
- Sex differences in fear memory consolidation via Tac2 signaling in mice
Techniques:
- 1p in vivo calcium imaging in freely moving mice.
- fNIRS and EEG during FPS and EEG in humans.
- Molecular biology.
- Machine learning.
Suitable background: Highly motivated individuals with relevant experience and strong publication records in the proposed topics.
Candidates, please contact Dr. Raül Andero: raul.andero@uab.cat
This project seeks to uncover the molecular mechanisms underlying selective cell vulnerability involved in neurodegeneration and dysfunction of neural circuits in Alzheimer’s disease. By profiling gene expression and metabolic alterations at single-cell resolution, the project aims to identify cell-type-specific metabolic pathways driving synaptic dysfunction, inflammation, and disease progression.
Our laboratory integrates innovative molecular strategies with state-of-the-art genomic, transcriptomic, proteomic, and metabolic approaches to define the brain cell types and functional states most susceptible to Alzheimer’s pathology. The successful candidate will employ cell-specific multi-omics and super-resolution imaging with lipid biosensors in human and mouse brains, combined with gene transfer, pharmacological and chemogenetic manipulations of memory and emotional memory circuits in advanced Alzheimer’s disease mouse models.
Applicants should have a strong background in neuroscience/neuropharmacology, with expertise in systems biology, metabolomics and super-resolution confocal imaging analyses. Experience in RNA sequencing, spatial transcriptomics, bioinformatics, lipidomics, in vivo stereotaxic injections, and behavioral studies in neurodegeneration mouse models is highly desirable.
Candidates, please contact Dr. Carles Saura: carlos.saura@uab.cat
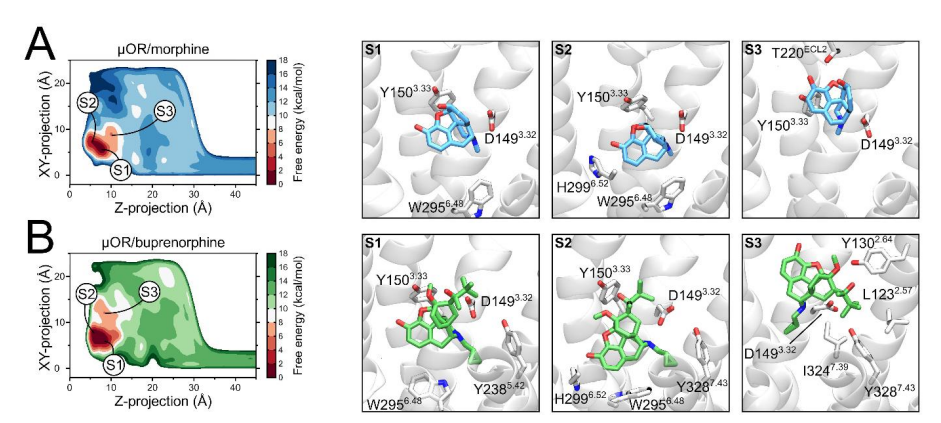
µ-opioid receptor (MOR) positive allosteric modulators that potentiate the action of endogenous opioids are promising drug candidates for the treatment of chronic pain with expected reduced side effects compared to marketed opioid drugs. However, the mechanism of MOR allosteric modulation is not well characterized, and the relationship between chemical structure and activity of MOR allosteric modulators is poorly understood. Computational methods can provide a mechanistic understanding of allosteric modulation of MOR in models of physiological and pathological conditions, which can benefit novel drug discovery. Our project proposal introduces further novelty in the use of computational methods to study modulation of MOR by including the transmembrane electrostatic potential and variations in the pH in the system models.MD simulations will be performed on systems including MOR, endogenous opioids and an allosteric modulator in different conditions of membrane electric field and pH. Mathematical modeling and machine learning techniques will be used to provide a connection between computational structural data and pharmacological properties. Examples of recent publications:
- Structural Determinants of Buprenorphine Partial Agonism at the μ-Opioid Receptor
- Time-dependent ligand-receptor binding kinetics and functionality in a heterodimeric receptor model
- Allosteric binding cooperativity in a kinetic context
- Structural Assessment of Agonist Efficacy in the μ-Opioid Receptor: Morphine and Fentanyl Elicit Different Activation Patterns
Techniques:
- Molecular dynamics simulations
- Metadynamics
- Ordinary differential equations
- Machine learning
Suitable background: Computational biophysics; Computational biochemistry, bioinformatics, mathematics.
Candidates, please contact Dr. Jesús Giraldo: jesus.giraldo@uab.cat
The project aims to uncover how altered GABAergic signaling contributes to cerebellar mitochondrial dysfunction and behavioral deficits in Rett syndrome. We focus on parvalbumin-expressing GABAergic neurons, which have high energetic demands and are particularly vulnerable in RTT, and on a newly identified mitochondrial pool of the GABAB receptor (mt-GABABR).To address these questions, we combine advanced microscopy, mitochondrial bioenergetics and proteomics, pharmacological modulation of GABABR, AAV-based genetic approaches, and cerebellar-dependent behavioral assays in RTT mouse models. This multidisciplinary approach will define the role of mt-GABABR in neuronal metabolism and evaluate its potential as a therapeutic target for RTT.
The candidate will have access to state-of-the-art approaches including expansion and super-resolution microscopy to resolve subcellular and mitochondrial protein localization, magnetic-activated cell sorting (MACS)–based isolation of neuronal subtype–specific mitochondria, mitochondrial bioenergetic analyses (including Seahorse extracellular flux measurements), cerebellar organotypic slices, quantitative mass spectrometry–based proteomics (LC–MS/MS), and AAV-mediated genetic manipulations. Pharmacological modulation of GABABR with arbaclofen, combined with cerebellar-dependent behavioral analyses in RTT mouse models, will enable a multiscale investigation of mitochondrial function spanning molecular, cellular, and behavioral levels.
Candidates should have a strong background in neuroscience, cell biology, or a related field, with a clear interest in neuronal metabolism, mitochondrial function, and neurodevelopmental disorders. Experience with advanced microscopy techniques (e.g., TEM, super-resolution microscopy) and in vivo mouse work is expected, while prior expertise in stereotaxic surgery, viral delivery, and behavioral assays will be highly valued. Familiarity with molecular, biochemical, or omics-based approaches is an advantage, though candidates with complementary skill sets and a strong motivation to develop new techniques are strongly encouraged to apply.
Candidates, please contact Dr. Laura Cutando: laura.cutando@uab.cat