Study on the transoceanic colonization of a Drosophila fly
Retrotransposons are mobile genes able to transpose (to move) by an intermediate RNA. They constitute the most abundant and widely distributed class of transposable elements in eukaryote genomes. Drosophila (the fruit fly) contains at least 23 different families representing 5-10% of the genome. A great amount of somatic (not inherited) and germinal (inherited) mutations results from insertions of transposable elements.
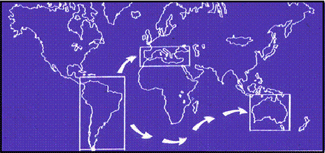
In this work, carried out by Dr María Pilar García Guerreiro and Dr Antonio Fontdevila from the Evolutionary Biology Group of the UAB, the molecular structure of the Osvaldo retrotransposon was studied in the D. buzzzatii species. This species originated in Argentina and colonized the Old World 300 years ago (Fig. 1).
Chromosomal distribution of this retrotransposon is different between original (from South America) and colonizing (European) populations. Original populations show a low occupancy frequency per chromosomal site compared to colonizing populations. In the latter mean occupancy frequency per site is higher because in addition to the low occupancy sites, several highly occupied sites are observed.
In order to explain this different distribution we have sequenced the DNA, in different colonizing populations, of three Osvaldo elements inserted in sites of high (2) and low (1) occupancy. We have observed that elements inserted in high insertion frequency sites show important nucleotide changes, duplications, and deletions, absent in the active element (with transposition ability). Moreover these sequences are identical and are located at the same nucleotide site in all colonizing populations (Fig. 2). On the contrary, the element inserted in the low occupancy site has the same sequence as the active element, except by a single nucleotide deletion.
These results favour the hypothesis that elements inserted in sites of high occupancy are inactive, their insertion antedated the colonization process and they arrived with the few founder individuals. Thus, most probably, their high insertion frequency is the result of the amplifying genetic drift effect of colonization by a small number of founders and not of an increase of the transposition rate. On the contrary, low occupancy sites host Osvaldo active copies (without nucleotide changes) that transposed after colonisation.

