Redesigning enzymes
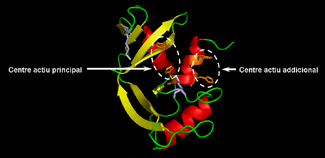
The chemical reactions that maintain the structure and function of living beings are catalyzed by enzymes. Enzymes are proteins that participate in most biological processes accelerating specific reactions. At the same time, and as opposed to conventional chemical catalyst, they can change the rate of a reaction according to the cell’s needs. Enzymes are highly specific both with respect to the substrates they recognize and to the products resulting from their action.
Enzymes are polymers in which the monomeric building blocks are amino acids; the specific amino acid sequence, as well as its three-dimensional structure -the placement of their atoms in space- is crucial for enzymatic activity. The region of the enzyme where the amino acids directly involved in the catalytic process are located is called the active site.
The enzymes showing ribonuclease (RNase) activity catalyze the breakdown of ribonucleic acids (RNAs); RNAs are polymers whose monomeric building blocks are ribonucleotides. RNase A is an enzyme with a very high catalytic efficiency. The group in the Department of Biochemistry and Molecular Biology (UAB) under the leadership of Dr. C.M. Cuchillo and Dr. M.V. Nogués has shown, with the use of different approaches that, in the case of RNase A, the catalytic process does not involve the specific interaction of the substrate with the active site of the enzyme, but also the correct positioning of the RNA polymer through non-catalytic binding sub-sites that are mostly located on the enzyme's surface.
Representation of the three-dimensional structure of RNase A and the predicted structures of modified forms incorporating a new active site.
The techniques called "site-directed mutagenesis" or "protein engineering" allow the selective modification of the amino acid sequence that specifies protein structure and function. The experimental work reported, carried out by Dr. M. Moussaoui, was based on these techniques and has allowed the preparation of modified forms of RNase A in which one of the binding sub-sites is transformed into a new catalytically active site. This has given rise to an enzyme with catalytic properties different from those of the native enzyme.
This work is a good example of how information obtained from basic research on the structure and function of enzymes can be the foundation of biotechnological applications aiming to the design of novel biocatalysts.
References
"A phosphate-binding subsite in bovine pancreatic ribonuclease A can be converted into a very efficient catalytic site". Mohammed Moussaoui, Claudi M. Cuchillo and M.Victòria Nogués. PROTEIN SCIENCE (2007) 16, 99-109.

