Medicine and Health
Development of a Harmless Artificial Virus for Gene Therapy
Researchers of the Nanobiology Unit from the UAB Institute of Biotechnology and Biomedicine, led by Antonio Villaverde, have produced an alternative to the use of viruses in gene therapy.The researchers synthesised nanoparticles which act as artificial viruses, capable of surrounding DNA fragments and releasing them as therapeutic agents, with no biological risk, into the interior of cells.
References
Unzueta, U.; Saccardo, P.; Domingo-Espín, J.; Cedano, J.; Conchillo-Solé, O.; García-Fruitós, E.; Céspedes, M.V.; Corchero, J.L.; Daura, X.; Mangues, R.; Ferrer-Miralles, N.; Villaverde, A.; Vázquez, E. Sheltering DNA in self-organizing, protein-only nano-shells as artificial viruses for gene delivery. Nanomedicine. 2014, vol. 10, num. 3, p. 535-41. doi: 10.1016/j.nano.2013.11.006.
Non-viral gene therapy, which is the insertion of genes into the genome with therapeutic aims without using viruses,and in general emerging nanomedicines aim to mimic virus through tuneable nanoparticles, for the cell targeted delivery of nucleic acids and other drugs.
Among a diversity of tested materials, proteins offer biocompatibility, biodegradability, and a wide spectrum of functionalities that can be adjusted by genetic engineering. Such a functional versatility is in contrast with the null control so far exercised over the organization of de novo designed building blocks to form protein complexes. While natural protein nanoparticles take advantage of the evolution to optimize the self-assembling, the novo multifunctional proteins fail to reach predefined supramolecular organization.
Recently, Professor Antonio Villaverde’s group has discovered the combination necessary to make these proteins act as an artificial virus and self-assemble themselves to form regular nanoparticles capable of penetrating target cells and reaching the nucleus in a very efficient manner. The key lies in a combination of cation-peptide and hexahistidine placed respectively at the amino and C-terminus ends of the modular proteins.
Although these protein nanoparticles efficiently bind plasmid DNA for subsequent expression of these transgenes and are very promising tools in nanomedicine, their supramolecular organization has been unexplored. In this article we have shown how, in the presence of DNA, there’s a reorientation of cationic segments on the inner surface that promote a supramolecular organization resulting in a protein closed coverage of inner nucleic acids in a virus-like manner,which results in DNA protection against external nucleases.
When R9-GFP-H6 proteins are in contact with foreign DNA, the protein forms spontaneous shells embedding DNA that reminds the organization of viral proteins, presenting both, isometric and rod-shaped architectonic models.
Interestingly, the capacity of the cationic end-terminal tags to promote protein self-assembling seems irrespective of the chosen core polypeptide. This allows the selection of homologous core proteins to form nanoparticles in order to avoid any immune response upon systemic administration, which could be a critical bottleneck to the therapeutic use.
In summary, we have shown for the first time how protein-based artificial viruses formed by self-assembling proteins shielding a DNA nucleus, can be generated by the fully de novo design of building blocks. This fact not only validates R9 and H6 peptides in vehicles for non-viral gene therapy, but also reveals an unexpected architectonic potential of these tags in the generation of tuneable protein shells, whose properties can be further adjusted by protein engineering. These versatile agents are promising alternatives to viruses since they overcome limitations such as rigid architecture and biosafety concerns.
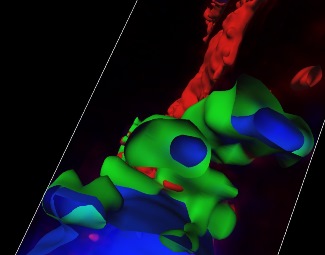
Top left figure. 3D reconstruction of serial confocal sections showing protein (green)-DNA (blue) nanoparticles.
Antonio Villaverde
Esther Vázquez
Department of Genetics and Microbiology
Institute of Biotechnology and Biomedicine (IBB)
2026 Universitat Autònoma de Barcelona
B.11870-2012 ISSN: 2014-6388
